The bottleneck in rare disease diagnostics has shifted. Next-Generation Sequencing provides unprecedented genomic data, but interpretation remains the challenge. In its latest white paper, SeqOne demonstrates how the new version of DiagAI, the variant prioritization engine within their CE-IVD NGS platform, delivers breakthrough automated genomic analysis validated against exceptionally rigorous clinical data.
Tested on 565 whole-genome sequencing cases from the Genomics England 100,000 Genomes Project, DiagAI Germline proves that high-precision automated diagnosis is now achievable at true population scale.
From Local to Large-Scale Representative Validation
Building a strong AI model is one step. Proving it works across the full diversity of real-world clinical cases is the definitive test.
Most variant prioritization tools have been validated on curated local cohorts. These are useful for development but limited in the breadth of diseases, sequencing protocols, and patient populations they represent. The Genomics England 100,000 Genomes Project is a fundamentally different proving ground: a national-scale, clinically diverse dataset spanning neurodevelopmental disorders, cardiac conditions, rare syndromic presentations, and more.
SeqOne submitted DiagAI Germline to a representative subset of this gold-standard reference. The results set a new standard for the field.
The Numbers That Matter
Across 565 solved WGS cases from Genomics England, DiagAI Germline delivered:
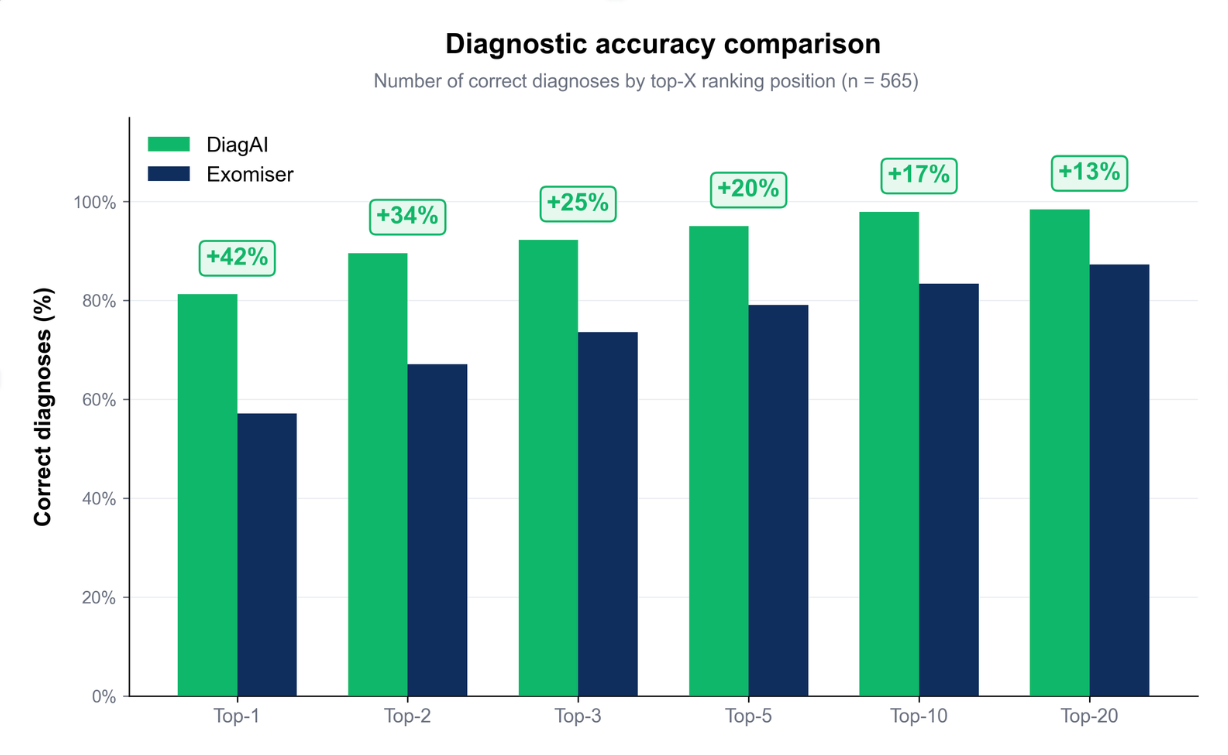
- 81% Top-1 Accuracy: The confirmed causal variant was ranked first in 81% of cases, consistently outperforming Exomiser, one of the most widely used prioritization tools in the field.
- 98% Clinical Inclusion: In 98% of cases, the diagnostic variant appeared within the top 10 results, ensuring near-complete coverage for clinical review.
- 96.6% SmartPick Precision: DiagAI's automated flagging system, SmartPick, achieved 96.6% precision, giving clinicians a high-confidence starting point for rapid case triage.
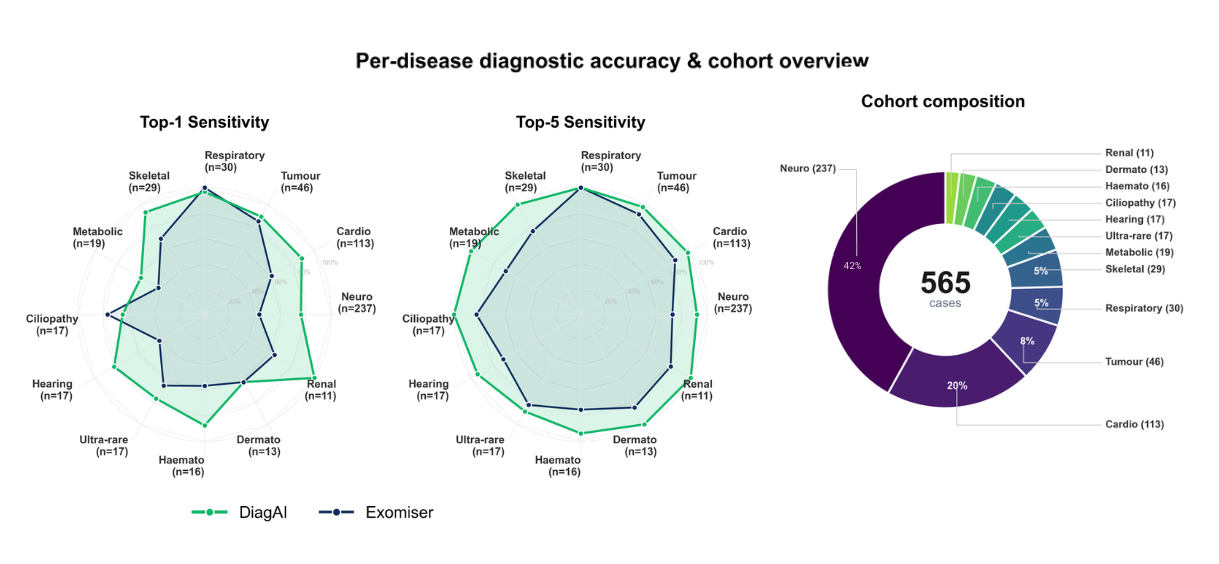
- Consistency Across Disease Areas: Performance held steady across diverse clinical categories, from neurological to cardiac to rare syndromic conditions. This is a model that generalizes.
These results are reinforced by long-term validation on 1,012 nephrology exomes, where DiagAI achieved an 84.2%Top-1 ranking. This confirms that the engine performs reliably across both exome and genome datasets and across independent clinical cohorts.
What Makes Diag AI Germline Different
At the heart of DiagAI Germline is UP², a unified scoring framework that evaluates every variant, regardless of type, on a single 0-to-100 scale. Rather than relying on a single signal, UP² integrates molecular pathogenicity predictions, phenotype-genotype alignment, inheritance compatibility, family segregation, and technical quality into one transparent score.
Clinicians do not need to mentally reconcile outputs from multiple disconnected tools. They receive a single, ranked shortlist, averaging just 6 to 8 variants per case, that reflects the full weight of available evidence.
Full Transparency, by Design
In a regulated clinical environment, a high score is only as valuable as the reasoning behind it. DiagAI is built on a "no black box" principle. Every prioritization decision is fully traceable through:
- A Summary Score Overview detailing the primary analytical components behind the DiagAI Score.
- A Pathogenicity Contribution Chart: a visual waterfall showing how specific genomic features drive the UP² molecular prediction for each variant.
- Phenotype-Genotype Alignment mapping HPO term matches between the patient's clinical profile and the gene's established phenotypic associations.
- A Clinical Evidence Trace logging every score adjustment (inheritance mode, segregation, quality filters, PanelApp status) so the clinician can see exactly why a variant was ranked where it was.
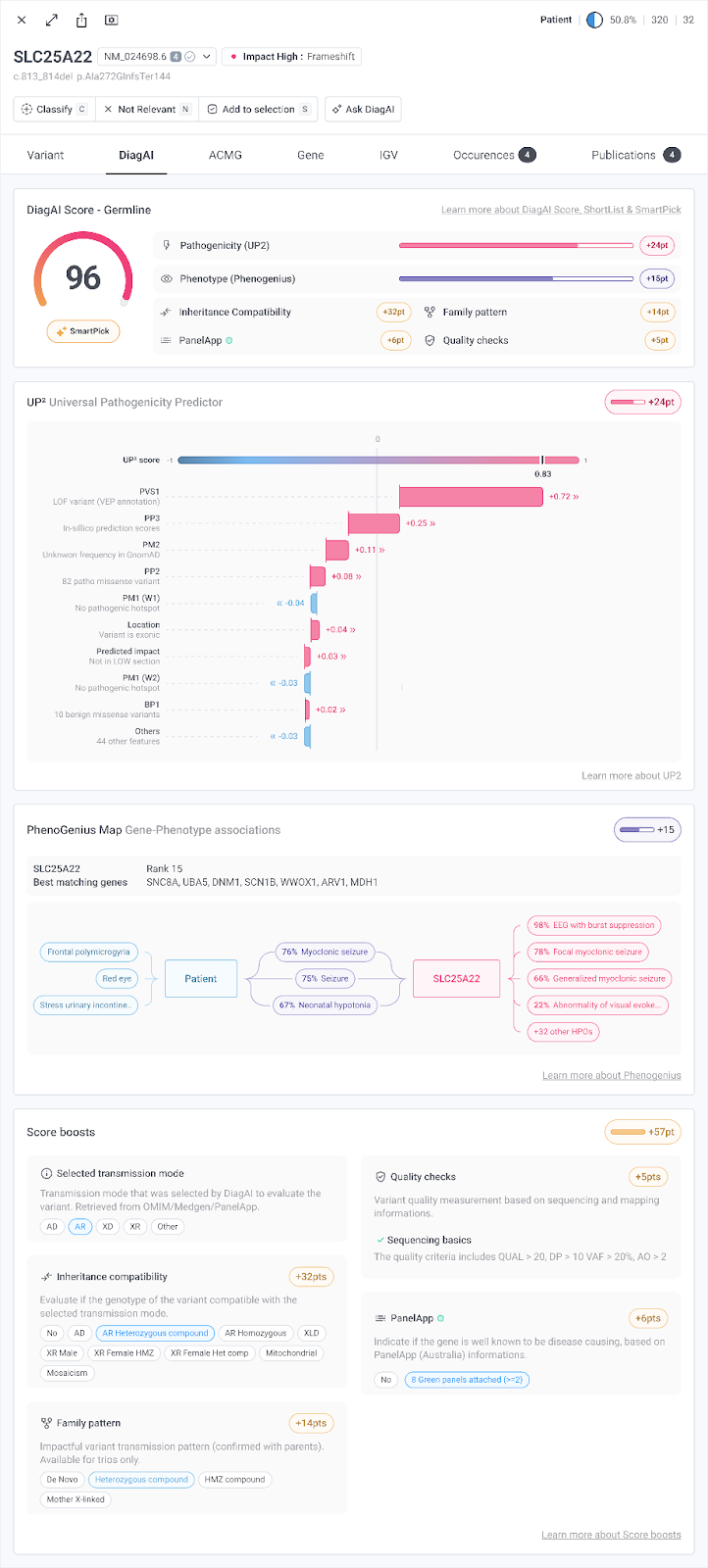
The clinician remains the final authority. DiagAI surfaces the most relevant variants with full evidence so that expert review is faster and more focused, not replaced.
Structural Variants, Now on the Same Scale
The Genomics England validation is the headline, but the whitepaper goes further. It also details DiagAI's latest advancement in structural variant prioritization: the new UP²-SV tool. This dedicated XGBoost classifier brings deletions and duplications onto the same unified scoring scale as point mutations, with a validated 97.6% recall on clinical benchmarks. For laboratories looking to close the structural variant gap, this is essential reading.
CE-IVD Certified: A Regulatory-Grade Platform for Clinical Genomics
DiagAI brings advanced intelligence to the clinical frontline via the SeqOne Platform, a CE-marked Class C in vitro diagnostic medical device under EU Regulation 2017/746 (IVDR). This certification means the platform has met the stringent safety, performance, and quality requirements demanded of clinical-grade diagnostic software in the European Union. For laboratories and health systems evaluating genomic tools, CE-IVD certification provides the assurance that DiagAI has been independently assessed for use in real clinical decision-making.
A New Standard for Genomic Medicine
DiagAI now resolves the total genomic burden of a patient, from SNVs to structural rearrangements, within a single, transparent, clinically validated, and CE-IVD certified framework. This is the convergence that genomic medicine has been working toward: comprehensive variant interpretation, proven at population scale, ready for the clinic.
Download the whitepaper to explore the full validation data, methodology, and UP²-SV details.





























